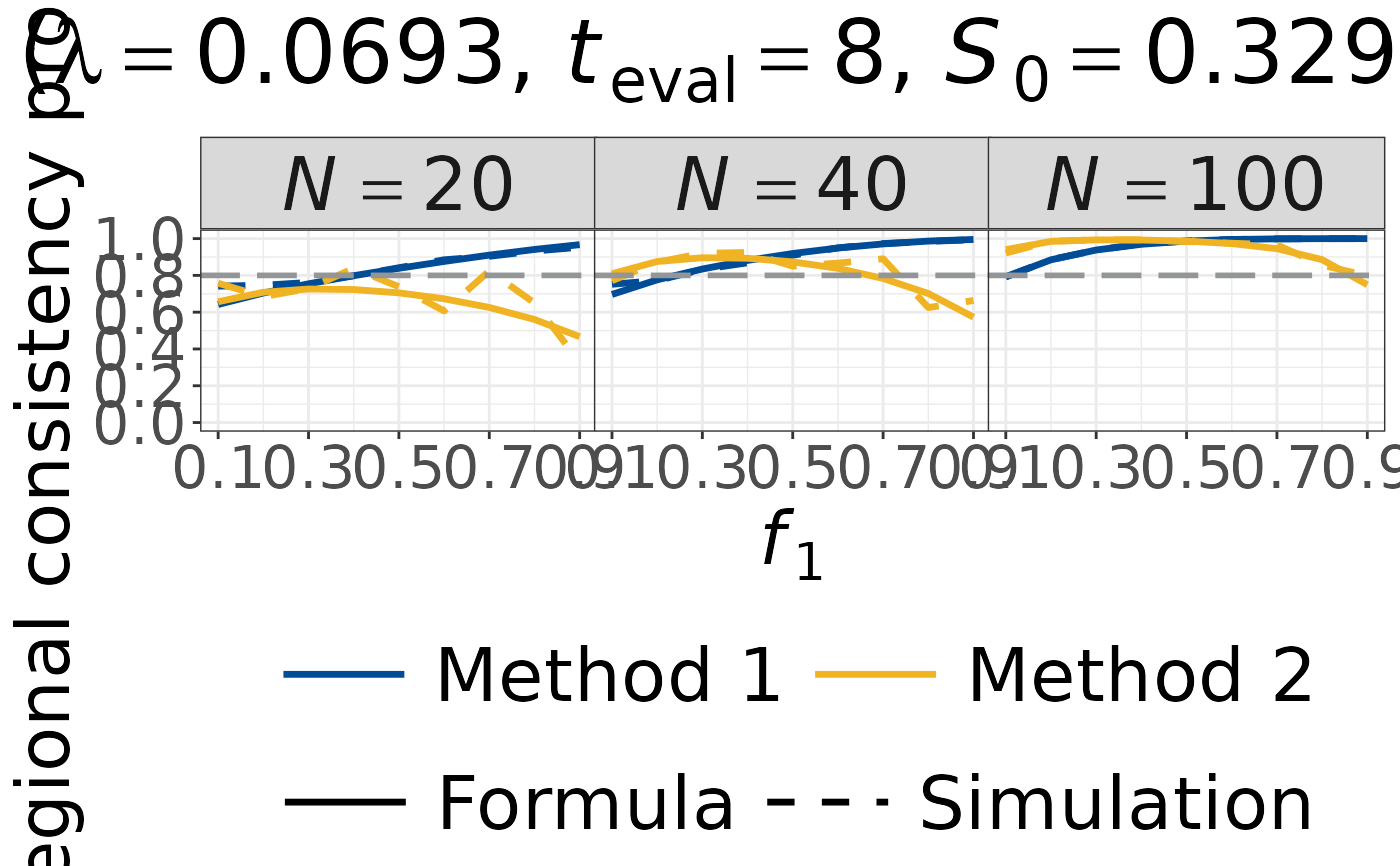
Plot Regional Consistency Probability for Single-Arm MRCT (Milestone Survival Endpoint)
Source:R/plot_rcp1armMilestoneSurvival.R
plot_rcp1armMilestoneSurvival.RdGenerate a faceted plot of Regional Consistency Probability (RCP) as a function
of the regional allocation proportion \(f_1\) for milestone survival endpoints.
Formula and simulation results are shown together for both Method 1 and Method 2.
Facet columns correspond to total sample sizes specified in N_vec.
Regional sample sizes are allocated as: \(N_{j1} = \lfloor N \times f_1 \rfloor\) and \(N_{j2} = \cdots = N_{jJ} = (N - N_{j1}) / (J - 1)\).
Arguments
- lambda
Numeric scalar. True hazard rate under the alternative hypothesis. Must be positive. Default is
log(2) / 10.- t_eval
Numeric scalar. Milestone evaluation time point. Must be positive. Default is
8.- S0
Numeric scalar. Historical control survival rate at
t_eval. Must be in \((0, 1)\). Default isexp(-log(2) * 8 / 5).- t_a
Numeric scalar. Accrual period. Must be positive. Default is
3.- t_f
Numeric scalar. Follow-up period. Must be positive. Default is
10.- lambda_dropout
Numeric scalar or
NULL. Dropout hazard rate. IfNULL(default), no dropout is assumed.- PI
Numeric scalar. Effect retention threshold for Method 1. Must be in \([0, 1]\). Default is
0.5.- N_vec
Integer vector. Total sample sizes for each facet column. Default is
c(20, 40, 100).- J
Positive integer (>= 2). Number of regions. Default is
3.- f1_seq
Numeric vector. Sequence of Region 1 allocation proportions. Each value must be in \((0, 1)\). Default is
seq(0.1, 0.9, by = 0.1).- nsim
Positive integer. Number of Monte Carlo iterations for simulation. Default is
10000.- seed
Non-negative integer. Random seed for simulation. Default is
1.- base_size
Positive numeric. Base font size in points passed to
theme. Use larger values (e.g.,28) for presentation slides and smaller values (e.g.,11) for vignettes or reports. Default is28.