
Survival Endpoints: Hazard Ratio, Milestone Survival, and RMST
Source:vignettes/survival-endpoints.Rmd
survival-endpoints.RmdThis vignette describes Regional Consistency Probability (RCP) calculations for three survival endpoint types: hazard ratio, milestone survival probability, and restricted mean survival time (RMST). All three endpoints share a common trial design framework, and event times are modelled by the exponential distribution.
Common trial design framework
For all survival endpoints, the following parameters define the trial design.
| Parameter | Symbol | Description |
|---|---|---|
t_a |
Accrual period; patients enrol uniformly over | |
t_f |
Follow-up period after accrual closes | |
tau |
Total study duration (computed internally) | |
lambda |
True hazard rate under the alternative (exponential model) | |
lambda0 |
Historical control hazard rate | |
lambda_dropout |
Dropout hazard rate; NULL assumes no dropout
(default) |
The common parameters below are used throughout the examples in this vignette.
lambda <- log(2) / 10 # treatment arm: median survival = 10
lambda0 <- log(2) / 5 # historical control: median survival = 5
t_a <- 3 # accrual period
t_f <- 10 # follow-up period
# True HR = lambda / lambda0 = 0.51. Hazard Ratio Endpoint
Statistical model
Under the exponential model with uniform accrual over and administrative censoring at , the expected event probability per patient is:
The expected number of events in Region is , and the log-hazard ratio estimator has the approximate distribution:
Consistency criteria
Method 1 (log-HR scale):
Letting , , and :
Method 1 (linear-HR scale) (Hayashi and Itoh 2018):
This is derived via the delta method. Define . Under homogeneity:
and .
Method 2:
Example
result_f <- rcp1armHazardRatio(
lambda = lambda,
lambda0 = lambda0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Time-to-Event (Hazard Ratio)
#>
#> Approach : Closed-Form Solution
#> True Hazard : lambda = 0.069315
#> Control Hazard : lambda0 = 0.138629
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall):
#> Log-HR based : 0.8935
#> Linear-HR based : 0.9228
#> Method 2 (All Regions Show Benefit):
#> HR < 1 : 0.9892
result_s <- rcp1armHazardRatio(
lambda = lambda,
lambda0 = lambda0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Time-to-Event (Hazard Ratio)
#>
#> Approach : Simulation-Based (nsim = 10000)
#> True Hazard : lambda = 0.069315
#> Control Hazard : lambda0 = 0.138629
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall):
#> Log-HR based : 0.9019
#> Linear-HR based : 0.9320
#> Method 2 (All Regions Show Benefit):
#> HR < 1 : 0.9924Effect of dropout
Specifying lambda_dropout reduces the expected event
probability
,
which in turn reduces all RCP values.
result_dropout <- rcp1armHazardRatio(
lambda = lambda,
lambda0 = lambda0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = 0.05,
PI = 0.5,
approach = "formula"
)
print(result_dropout)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Time-to-Event (Hazard Ratio)
#>
#> Approach : Closed-Form Solution
#> True Hazard : lambda = 0.069315
#> Control Hazard : lambda0 = 0.138629
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = 0.050000
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall):
#> Log-HR based : 0.8656
#> Linear-HR based : 0.8971
#> Method 2 (All Regions Show Benefit):
#> HR < 1 : 0.9793Visualisation
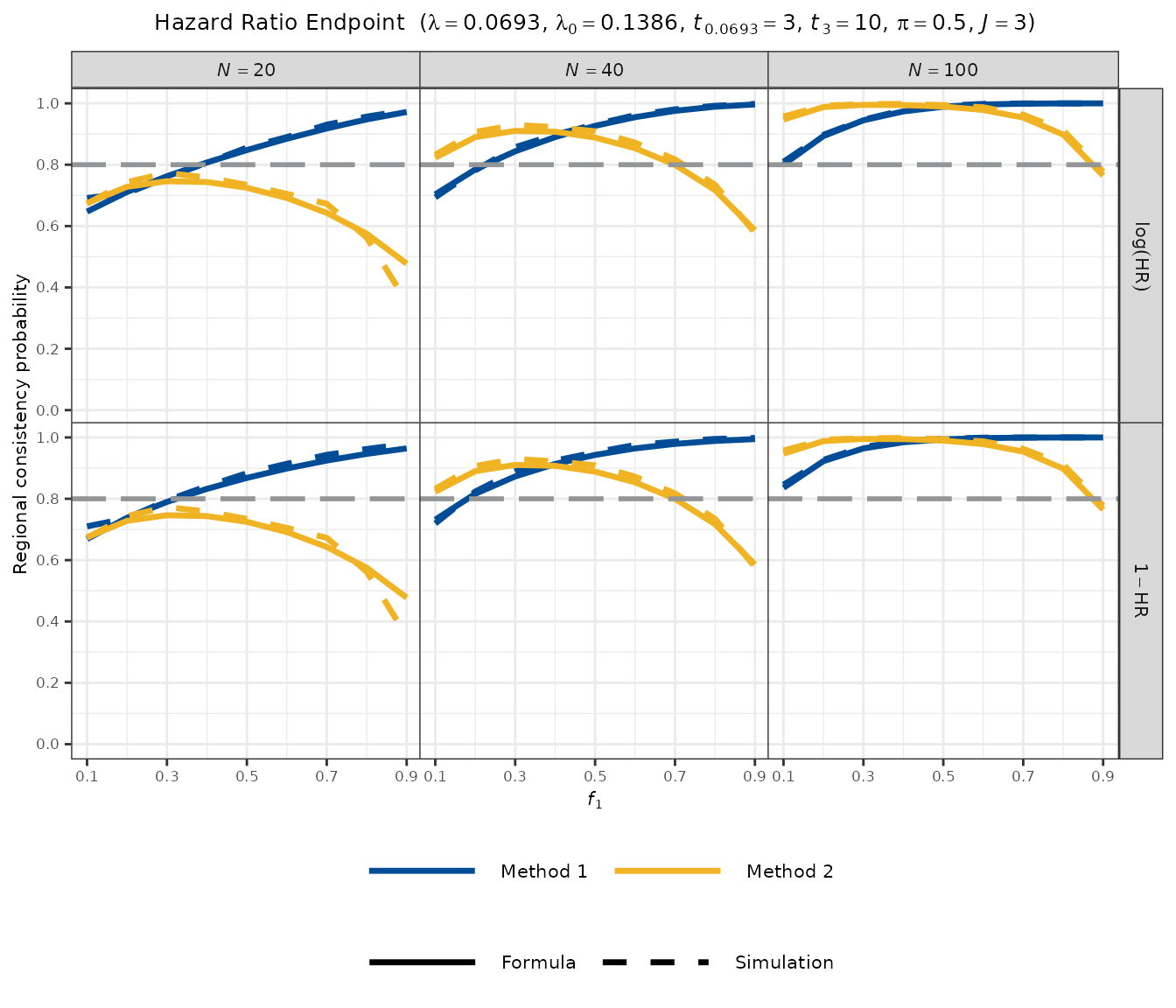
plot_rcp1armHazardRatio(
lambda = lambda,
lambda0 = lambda0,
t_a = t_a,
t_f = t_f,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 8
)
2. Milestone Survival Endpoint
Statistical model
The treatment effect at evaluation time is:
where is the historical control survival rate at , supplied by the user.
The asymptotic variance of the Kaplan-Meier estimator is derived from Greenwood’s formula:
where:
Closed-form solution when ( throughout):
For
this reduces to the binomial variance
.
When
,
the integral is evaluated numerically via
stats::integrate().
Example
t_eval <- 8
S0 <- exp(-log(2) * t_eval / 5)
cat(sprintf("True S(%g) = %.4f, S0 = %.4f, delta = %.4f\n",
t_eval, exp(-lambda * t_eval), S0,
exp(-lambda * t_eval) - S0))
#> True S(8) = 0.5743, S0 = 0.3299, delta = 0.2445
result_f <- rcp1armMilestoneSurvival(
lambda = lambda,
t_eval = t_eval,
S0 = S0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Milestone Survival
#>
#> Approach : Closed-Form Solution (Greenwood)
#> True Hazard : lambda = 0.069315
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#> Eval Time : t_eval = 8.00
#> Control Surv : S0 = 0.3299
#> True Surv : S_est = 0.5743
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8848
#> Method 2 (All Regions > S0) : 0.9865Since
,
the closed-form solution is used
(formula_type = "closed-form"). For
,
numerical integration is applied automatically.
result_s <- rcp1armMilestoneSurvival(
lambda = lambda,
t_eval = t_eval,
S0 = S0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Milestone Survival
#>
#> Approach : Simulation-Based (nsim = 10000)
#> True Hazard : lambda = 0.069315
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#> Eval Time : t_eval = 8.00
#> Control Surv : S0 = 0.3299
#> True Surv : S_est = 0.5743
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8873
#> Method 2 (All Regions > S0) : 0.9887Visualisation
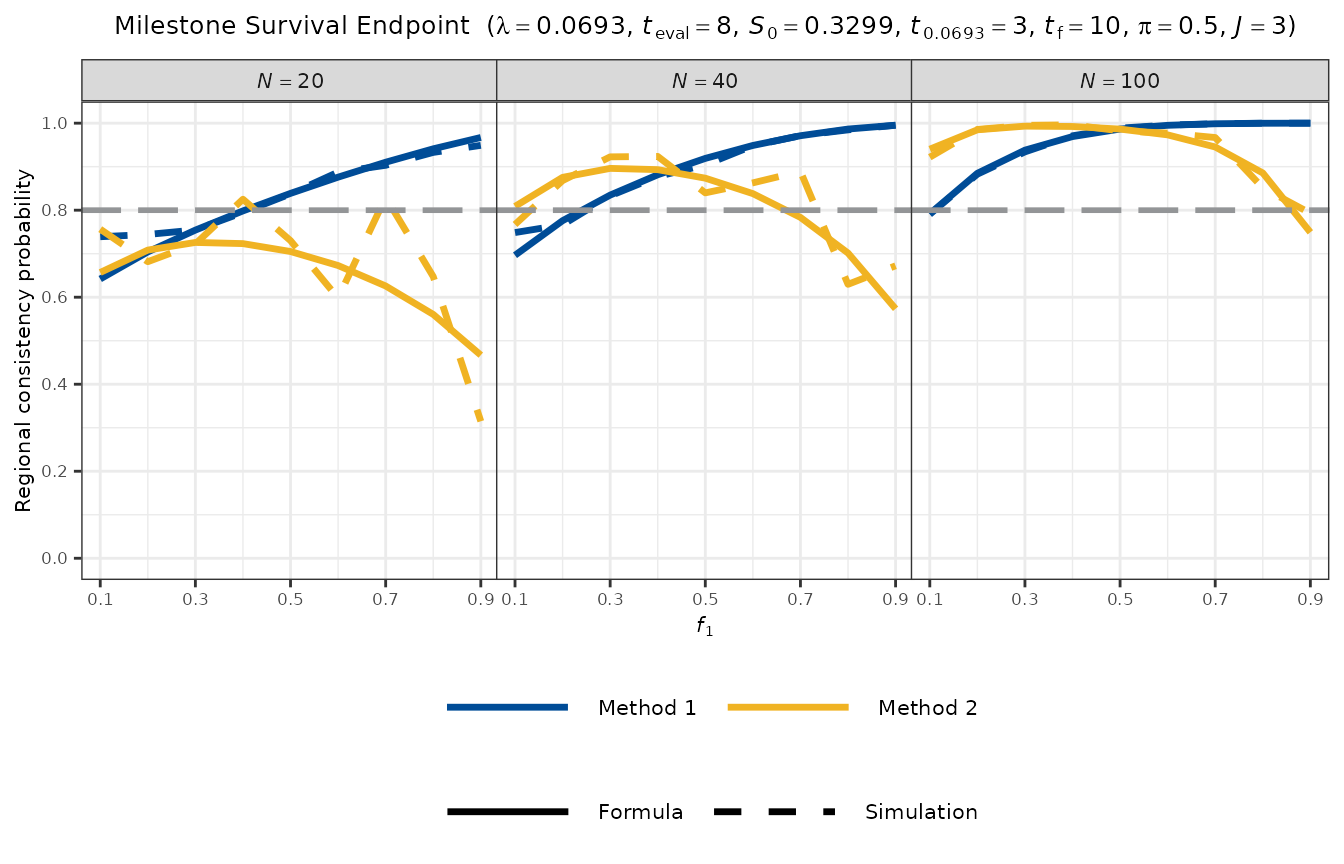
plot_rcp1armMilestoneSurvival(
lambda = lambda,
t_eval = t_eval,
S0 = S0,
t_a = t_a,
t_f = t_f,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 8
)
3. RMST Endpoint
Statistical model
The restricted mean survival time (RMST) up to truncation time is:
The asymptotic variance of the regional RMST estimator is:
where:
Closed-form solution when , using (and ):
$$ v_{\text{RMST}}(\tau^*) = \frac{1}{\lambda}\Bigl[ A(\lambda_d) - 2\,e^{-\lambda\tau^*}\,A(\lambda_d + \lambda) + e^{-2\lambda\tau^*}\,A(\lambda_d + 2\lambda) \Bigr] $$
For this simplifies to:
When
,
the integral is split at
and evaluated via stats::integrate().
Example
tau_star <- 8
mu0 <- (1 - exp(-lambda0 * tau_star)) / lambda0
mu_est <- (1 - exp(-lambda * tau_star)) / lambda
cat(sprintf("True RMST = %.4f, mu0 = %.4f, delta = %.4f\n",
mu_est, mu0, mu_est - mu0))
#> True RMST = 6.1408, mu0 = 4.8339, delta = 1.3069
result_f <- rcp1armRMST(
lambda = lambda,
tau_star = tau_star,
mu0 = mu0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Restricted Mean Survival Time (RMST)
#>
#> Approach : Closed-Form Solution
#> True Hazard : lambda = 0.069315
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#> Trunc. Time : tau* = 8.00
#> Control RMST : mu0 = 4.8339
#> True RMST : mu_est = 6.1408
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8693
#> Method 2 (All Regions > mu0) : 0.9808
result_s <- rcp1armRMST(
lambda = lambda,
tau_star = tau_star,
mu0 = mu0,
Nj = c(20, 80),
t_a = t_a,
t_f = t_f,
lambda_dropout = NULL,
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Restricted Mean Survival Time (RMST)
#>
#> Approach : Simulation-Based (nsim = 10000)
#> True Hazard : lambda = 0.069315
#> Sample Size : Nj = (20, 80)
#> Total Size : N = 100
#> Accrual Period : t_a = 3.00
#> Follow-up : t_f = 10.00
#> Study Duration : tau = 13.00
#> Dropout Hazard : lambda_d = NA
#> Threshold : PI = 0.5000
#> Trunc. Time : tau* = 8.00
#> Control RMST : mu0 = 4.8339
#> True RMST : mu_est = 6.1408
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8790
#> Method 2 (All Regions > mu0) : 0.9809Visualisation
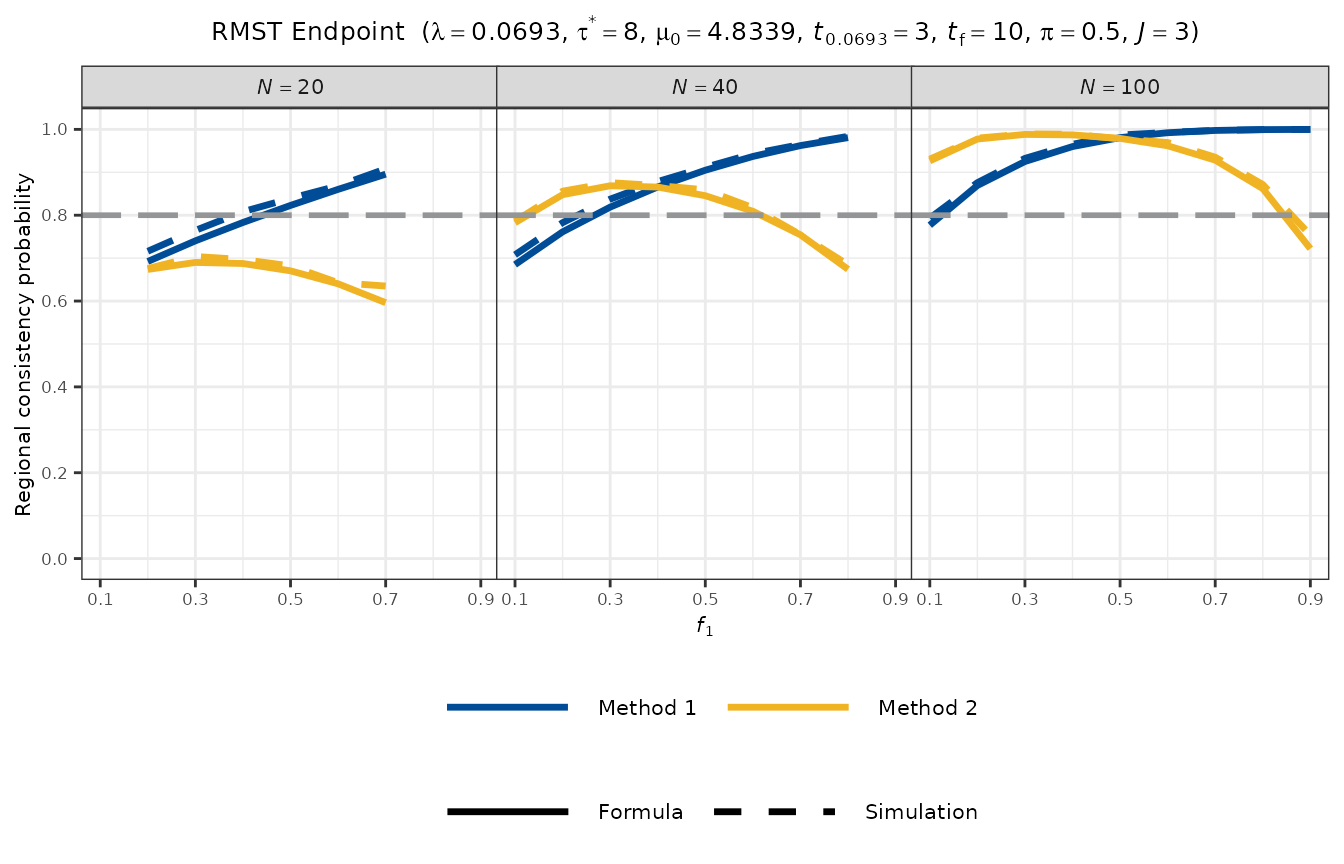
plot_rcp1armRMST(
lambda = lambda,
tau_star = tau_star,
mu0 = mu0,
t_a = t_a,
t_f = t_f,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 8
)
Summary
| Endpoint | Effect parameter | Benefit direction | Variance basis | Closed-form condition |
|---|---|---|---|---|
| Hazard Ratio | (Method 1, log-HR scale); (Method 1, linear-HR scale) | Expected events via (Wu 2015) | Always | |
| Milestone Survival | Greenwood’s formula | |||
| RMST | Squared survival difference integral |
References
Hayashi N, Itoh Y (2017). A re-examination of Japanese sample size calculation for multi-regional clinical trial evaluating survival endpoint. Japanese Journal of Biometrics, 38(2): 79–92. https://doi.org/10.5691/jjb.38.79
Wu J (2015). Sample size calculation for the one-sample log-rank test. Pharmaceutical Statistics, 14(1): 26–33. https://doi.org/10.1002/pst.1654