
Non-Survival Endpoints: Continuous, Binary, and Count
Source:vignettes/non-survival-endpoints.Rmd
non-survival-endpoints.RmdThis vignette describes Regional Consistency Probability (RCP) calculations for three non-survival endpoint types: continuous, binary, and count (negative binomial). For each endpoint, the statistical model, treatment effect scale, closed-form formulae, and worked examples are provided.
1. Continuous Endpoint
Statistical model
Let denote the sample mean for Region . Under the assumption that individual observations are independently and identically distributed as within each region, the regional sample means are:
independently across regions. The treatment effect relative to a historical control mean is .
Consistency criteria
Method 1 (Effect Retention):
Defining , the condition is equivalent to:
where is the sample mean pooled over regions . Under homogeneity:
Therefore:
Method 2 (Simultaneous Positivity):
Example
Setting: , , , ( regions with ), .
result_f <- rcp1armContinuous(
mu = 0.5,
mu0 = 0.1,
sd = 1,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Continuous
#>
#> Approach : Closed-Form Solution
#> Target Mean : mu = 0.5000
#> Null Mean : mu0 = 0.1000
#> Std. Dev. : sd = 1.0000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8340
#> Method 2 (All Regions > mu0) : 0.9522
result_s <- rcp1armContinuous(
mu = 0.5,
mu0 = 0.1,
sd = 1,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Continuous
#>
#> Approach : Simulation-Based (nsim = 10000)
#> Target Mean : mu = 0.5000
#> Null Mean : mu0 = 0.1000
#> Std. Dev. : sd = 1.0000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.8338
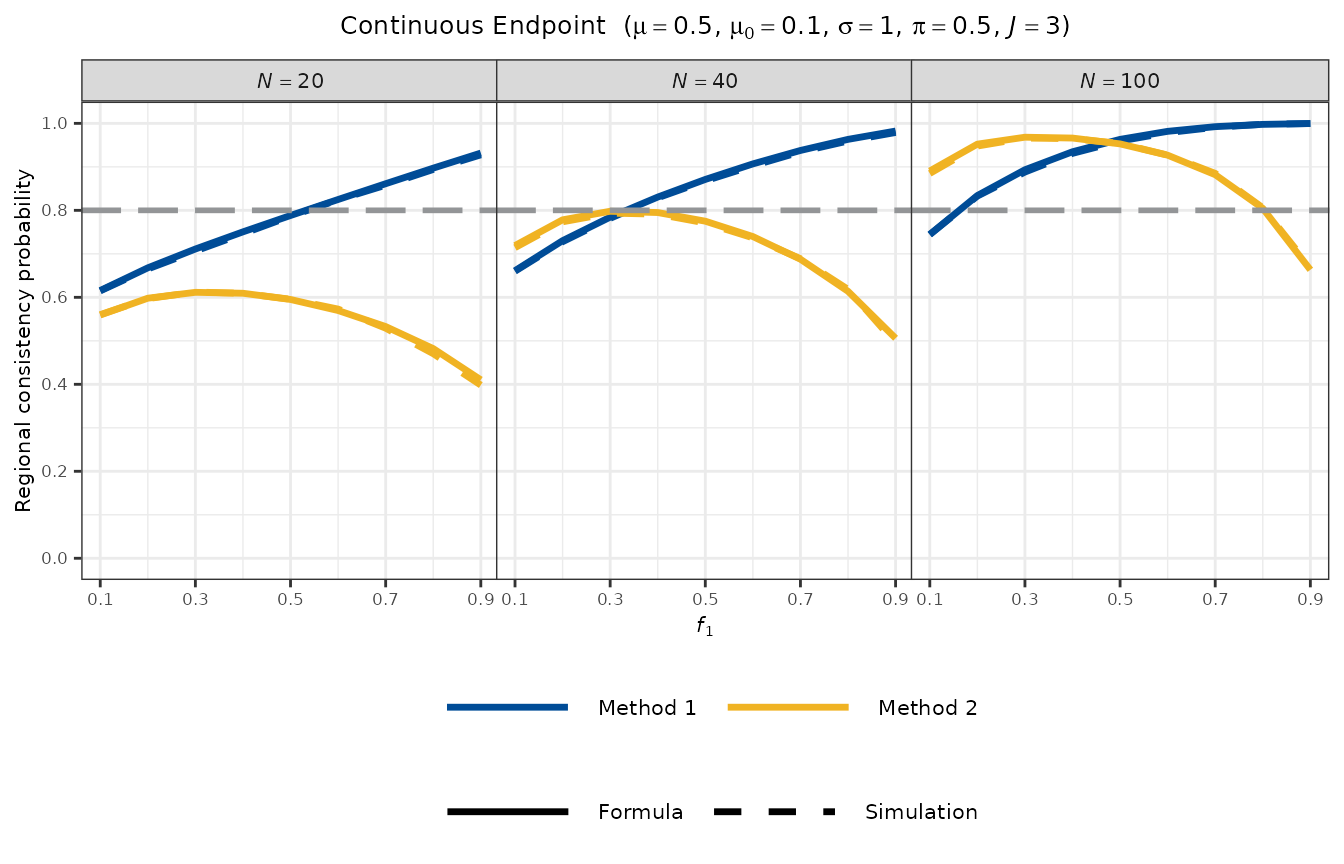
#> Method 2 (All Regions > mu0) : 0.9479Visualisation
plot_rcp1armContinuous(
mu = 0.5,
mu0 = 0.1,
sd = 1,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 8
)
2. Binary Endpoint
Statistical model
Let denote the number of responders in Region . Under independent Bernoulli trials with a common response rate :
independently across regions. The regional response rate estimator is , the overall estimator is , and the treatment effect is .
Consistency criteria
Method 1 (Effect Retention) — Exact Enumeration:
By the additivity of independent binomials, . The formula approach enumerates all combinations and sums the joint probabilities satisfying the consistency condition:
where .
Method 2 (Simultaneous Positivity):
The condition is equivalent to where . Denoting by the CDF of the binomial distribution with parameters and evaluated at :
Example
Setting: , , ( regions with ), .
result_f <- rcp1armBinary(
p = 0.5,
p0 = 0.2,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Binary
#>
#> Approach : Exact Solution
#> Response Rate : p = 0.5000
#> Null Rate : p0 = 0.2000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.9234
#> Method 2 (All Regions > p0) : 0.9939
result_s <- rcp1armBinary(
p = 0.5,
p0 = 0.2,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Binary
#>
#> Approach : Simulation-Based (nsim = 10000)
#> Response Rate : p = 0.5000
#> Null Rate : p0 = 0.2000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall) : 0.9203
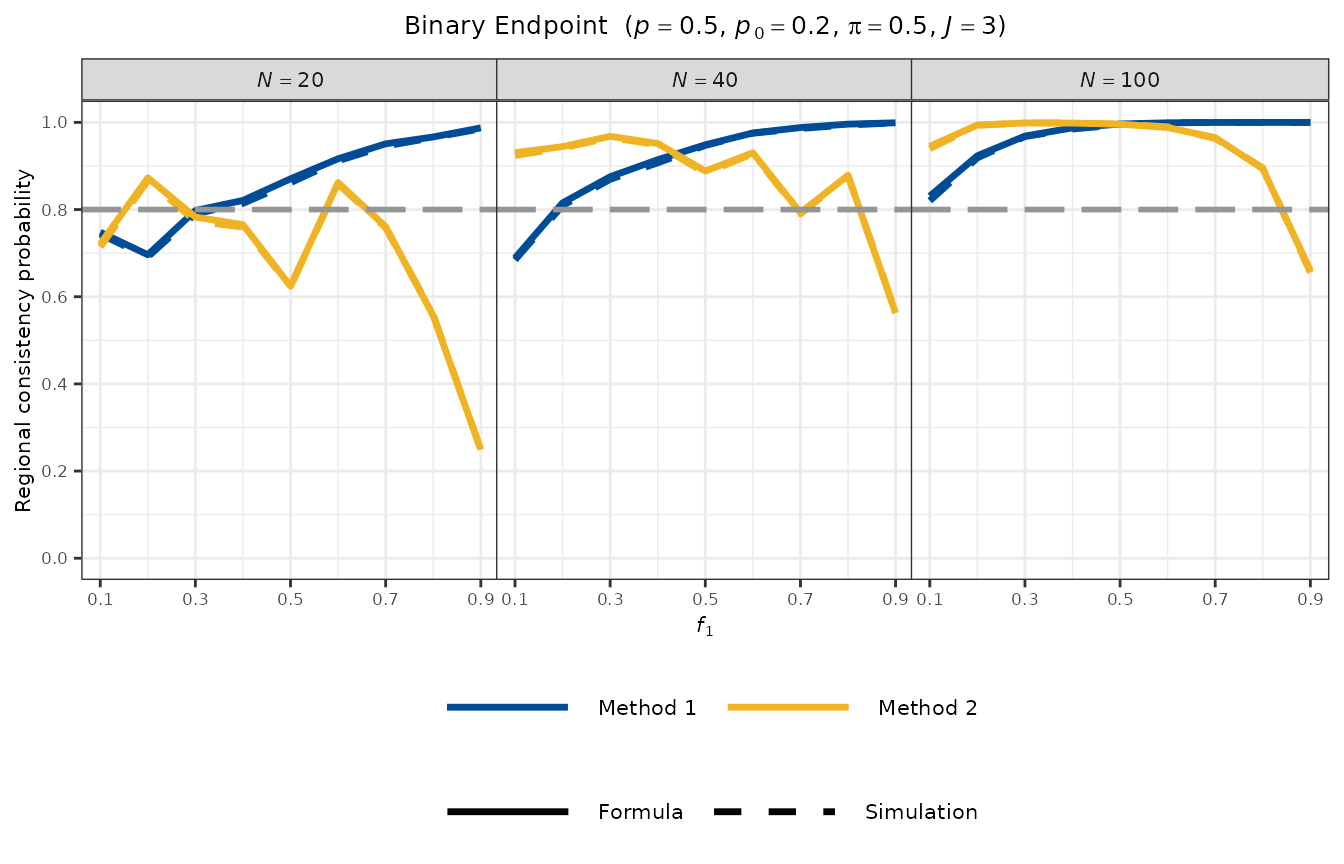
#> Method 2 (All Regions > p0) : 0.9933Visualisation
plot_rcp1armBinary(
p = 0.5,
p0 = 0.2,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 8
)
3. Count Endpoint (Negative Binomial)
Statistical model
Count data are modelled by the negative binomial distribution. The total event count in Region is:
independently across regions, where is the expected count per patient under the alternative and is the dispersion parameter. The regional rate estimator is , and the treatment effect is expressed as a rate ratio:
Benefit is indicated by .
By the reproducibility property of the negative binomial, the pooled count for regions follows , enabling exact enumeration analogous to the binary case.
Consistency criteria
Method 1 (log-RR scale):
Since (benefit), , so the condition requires to be sufficiently negative relative to the overall .
Method 1 (linear-RR scale):
Both Method 1 variants use exact enumeration over all combinations via the outer product of negative binomial PMFs.
Method 2:
Denoting by the CDF of the negative binomial distribution with mean and size evaluated at , the condition is equivalent to , i.e., when is not an integer (and otherwise). Therefore:
Example
Setting: , , , ( regions with ), .
result_f <- rcp1armCount(
lambda = 2,
lambda0 = 3,
dispersion = 1,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "formula"
)
print(result_f)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Count (Negative Binomial)
#>
#> Approach : Exact Solution
#> Expected Count : lambda = 2.000000
#> Control Count : lambda0 = 3.000000
#> Dispersion : dispersion = 1.000000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall):
#> Log-RR based : 0.8186
#> Linear-RR based : 0.8406
#> Method 2 (All Regions Show Benefit):
#> RR < 1 : 0.9320
result_s <- rcp1armCount(
lambda = 2,
lambda0 = 3,
dispersion = 1,
Nj = c(20, 40, 40),
PI = 0.5,
approach = "simulation",
nsim = 10000,
seed = 1
)
print(result_s)
#>
#> Regional Consistency Probability for Single-Arm MRCT
#> Endpoint : Count (Negative Binomial)
#>
#> Approach : Simulation-Based (nsim = 10000)
#> Expected Count : lambda = 2.000000
#> Control Count : lambda0 = 3.000000
#> Dispersion : dispersion = 1.000000
#> Sample Size : Nj = (20, 40, 40)
#> Total Size : N = 100
#> Threshold : PI = 0.5000
#>
#> Consistency Probabilities:
#> Method 1 (Region 1 vs Overall):
#> Log-RR based : 0.8118
#> Linear-RR based : 0.8331
#> Method 2 (All Regions Show Benefit):
#> RR < 1 : 0.9276The output reports three RCP values: Method 1 on the log-RR scale
(Method1_logRR), Method 1 on the linear-RR scale
(Method1_linearRR), and Method 2
(Method2).
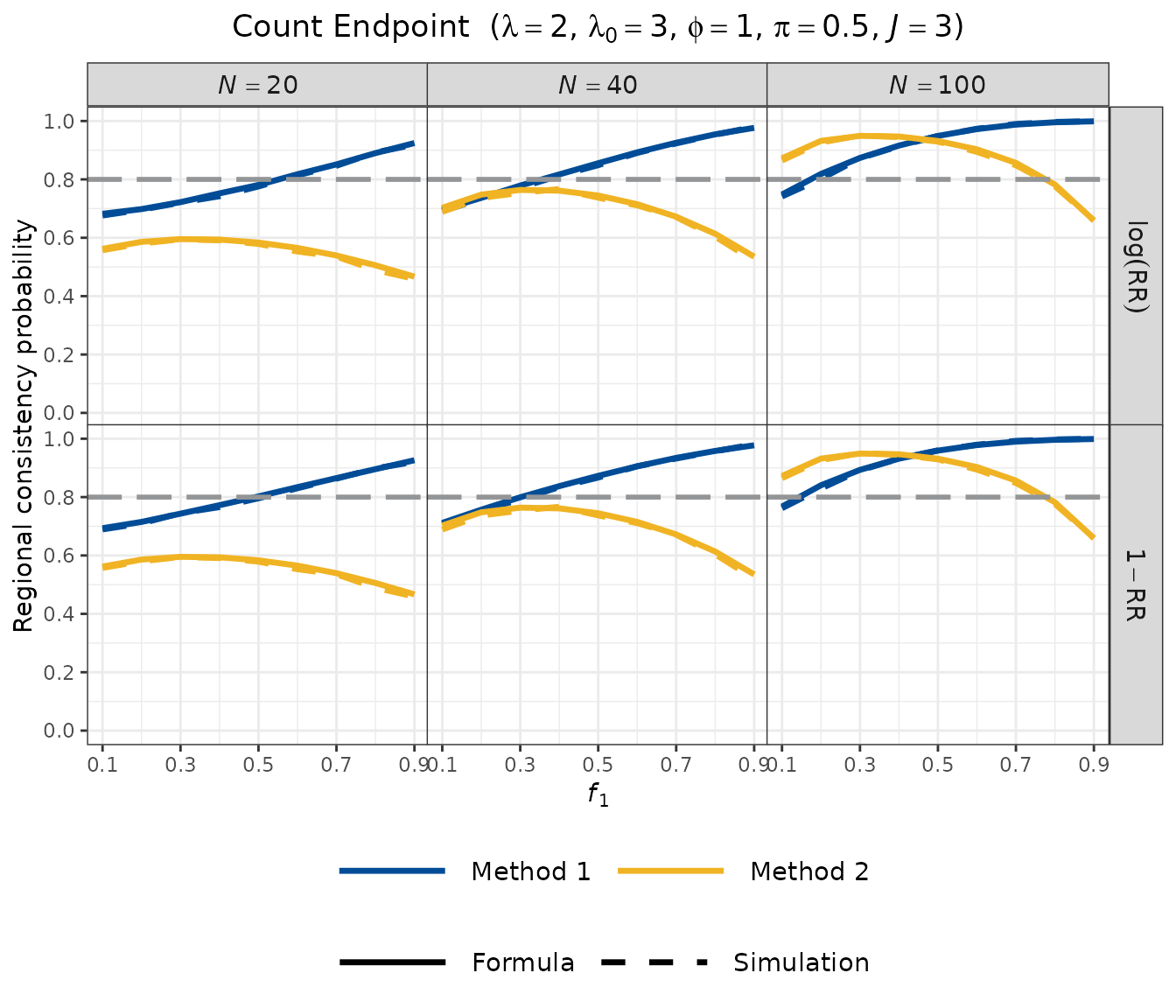
Visualisation
The count endpoint plot uses a grid layout: facet rows distinguish the two Method 1 scales (log-RR and ), and facet columns correspond to different total sample sizes.
plot_rcp1armCount(
lambda = 2,
lambda0 = 3,
dispersion = 1,
PI = 0.5,
N_vec = c(20, 40, 100),
J = 3,
nsim = 5000,
seed = 1,
base_size = 11
)
Summary
| Endpoint | Model | Effect parameter | Benefit direction | Method 1 computation | Method 2 computation |
|---|---|---|---|---|---|
| Continuous | Normal | Closed-form (normal approximation) | Product of normal tail probabilities | ||
| Binary | Binomial | Exact enumeration (binomial) | Product of binomial tail probabilities | ||
| Count | Negative binomial | (Method 1, log-RR scale); (Method 1, linear-RR scale) | Exact enumeration (negative binomial) | Product of NB tail probabilities |
References
Homma G (2024). Cautionary note on regional consistency evaluation in multiregional clinical trials with binary outcomes. Pharmaceutical Statistics, 23(3):385–398. https://doi.org/10.1002/pst.2358