Motivating Scenario
We extend the single-endpoint RA trial to a bivariate design with two co-primary continuous endpoints:
- Endpoint 1: change from baseline in DAS28 score
- Endpoint 2: change from baseline in HAQ-DI (Health Assessment Questionnaire – Disability Index)
The trial enrolls patients per group (treatment vs. placebo) in a 1:1 randomised controlled design.
Clinically meaningful thresholds (posterior probability):
| Endpoint 1 | Endpoint 2 | |
|---|---|---|
| 1.5 | 1.0 | |
| 0.5 | 0.3 |
Null hypothesis thresholds (predictive probability): , .
Decision thresholds: (Go), (NoGo).
Observed data (used in Sections 2–4):
- Treatment: ,
- Control: ,
where () is the sum-of-squares matrix.
1. Bayesian Model: Normal-Inverse-Wishart Conjugate
1.1 Prior Distribution
For group (: treatment, : control), we model the bivariate outcomes for patient () as
The conjugate prior is the Normal-Inverse-Wishart (NIW) distribution:
where
is the prior mean (argument mu0_t or mu0_c),
is the prior concentration (kappa0_t or
kappa0_c),
is the prior degrees of freedom (nu0_t or
nu0_c), and
is a
positive-definite prior scale matrix (Lambda0_t or
Lambda0_c).
The vague (Jeffreys) prior is obtained by letting
and
(prior = 'vague').
1.2 Posterior Distribution
Given
observations with sample mean
(ybar_t or ybar_c) and sum-of-squares matrix
(S_t or S_c), the NIW posterior has updated
parameters:
The marginal posterior of the group mean is a bivariate non-standardised -distribution:
Under the vague prior, this simplifies to
1.3 Posterior of the Bivariate Treatment Effect
The bivariate treatment effect is . Since the two groups are independent, the posterior of is the distribution of the difference of two independent bivariate -vectors:
where the posterior scale matrix is .
The posterior probability that falls in a rectangular region defined by thresholds on each component is
computed by one of two methods (MC or MM) described in Section 3.
1.4 Nine-Region Grid (Posterior Probability)
The bivariate threshold grid for
creates a
partition using theta_TV1, theta_MAV1,
theta_TV2, theta_MAV2:
| Endpoint 1 | ||||
|---|---|---|---|---|
| θ1 > θTV1 | θTV1 ≥ θ1 > θMAV1 | θMAV1 ≥ θ1 | ||
| Endpoint 2 | θ2 > θTV2 | R1 | R4 | R7 |
| θTV2 ≥ θ2 > θMAV2 | R2 | R5 | R8 | |
| θMAV2 ≥ θ2 | R3 | R6 | R9 | |
Region (both endpoints exceed ) is typically designated as Go and (both endpoints below ) as NoGo.
1.5 Four-Region Grid (Predictive Probability)
For the predictive probability, a
partition is used, defined by theta_NULL1 and
theta_NULL2:
| Endpoint 1 | |||
|---|---|---|---|
| θ1 > θNULL1 | θ1 ≤ θNULL1 | ||
| Endpoint 2 | θ2 > θNULL2 | R1 | R3 |
| θ2 ≤ θNULL2 | R2 | R4 | |
Region (both endpoints exceed ) is typically designated as Go and (both endpoints at or below ) as NoGo.
2. Posterior Predictive Distribution
The predictive distribution of the future sample mean
based on
future patients (m_t or m_c) is again a
bivariate
-distribution:
under the NIW prior. Under the vague prior the scale inflates to .
3. Two Computation Methods
The computation method is specified via the CalcMethod
argument.
3.1 Monte Carlo Simulation (CalcMethod = 'MC')
For each of
draws (nMC), independent samples are generated:
and the probability of region is estimated as the empirical proportion:
When pbayespostpred2cont() is called in vectorised mode
over
replicates, all
standard normal and chi-squared draws are pre-generated once and reused;
only the Cholesky factor of the replicate-specific scale matrix is
recomputed per observation.
3.2 Moment-Matching Approximation
(CalcMethod = 'MM')
The difference is approximated by a bivariate non-standardised -distribution (Yamaguchi et al., 2025, Theorem 3). Let be the scale matrix of (), and define
with . Then , where
The joint probability over each rectangular region is evaluated by a
single call to mvtnorm::pmvt(), which avoids simulation
entirely and is exact in the normal limit
().
The MM method requires for both groups ( in the notation above); if this condition is not met, the function falls back to MC automatically.
4. Study Designs
4.1 Controlled Design
Both groups are observed in the PoC trial. All NIW posterior parameters are updated from the current data.
Posterior probability – vague prior:
S_t <- matrix(c(18.0, 3.6, 3.6, 9.0), 2, 2)
S_c <- matrix(c(16.0, 2.8, 2.8, 8.5), 2, 2)
set.seed(42)
p_post_vague <- pbayespostpred2cont(
prob = 'posterior', design = 'controlled', prior = 'vague',
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = 20L,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = c(1.8, 1.0), S_c = S_c,
m_t = NULL, m_c = NULL,
kappa0_t = NULL, nu0_t = NULL, mu0_t = NULL, Lambda0_t = NULL,
kappa0_c = NULL, nu0_c = NULL, mu0_c = NULL, Lambda0_c = NULL,
r = NULL,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 5000L
)
print(round(p_post_vague, 4))
#> R1 R2 R3 R4 R5 R6 R7 R8 R9
#> 0.5056 0.2198 0.0000 0.1478 0.1264 0.0002 0.0000 0.0002 0.0000
cat(sprintf(
"g_Go = P(R1 | data) = %.4f\n", p_post_vague["R1"]
))
#> g_Go = P(R1 | data) = 0.5056
cat(sprintf(
"g_NoGo = P(R9 | data) = %.4f\n\n", p_post_vague["R9"]
))
#> g_NoGo = P(R9 | data) = 0.0000
cat(sprintf(
"Go criterion: g_Go >= gamma1 (0.80)? %s\n",
ifelse(p_post_vague["R1"] >= 0.80, "YES", "NO")
))
#> Go criterion: g_Go >= gamma1 (0.80)? NO
cat(sprintf(
"NoGo criterion: g_NoGo >= gamma2 (0.20)? %s\n",
ifelse(p_post_vague["R9"] >= 0.20, "YES", "NO")
))
#> NoGo criterion: g_NoGo >= gamma2 (0.20)? NO
cat(sprintf("Decision: %s\n",
ifelse(p_post_vague["R1"] >= 0.80 & p_post_vague["R9"] < 0.20, "Go",
ifelse(p_post_vague["R1"] < 0.80 & p_post_vague["R9"] >= 0.20, "NoGo",
ifelse(p_post_vague["R1"] >= 0.80 & p_post_vague["R9"] >= 0.20, "Miss",
"Gray")))
))
#> Decision: GrayPosterior probability – NIW informative prior:
L0 <- matrix(c(8.0, 0.0, 0.0, 2.0), 2, 2)
set.seed(42)
p_post_niw <- pbayespostpred2cont(
prob = 'posterior', design = 'controlled', prior = 'N-Inv-Wishart',
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = 20L,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = c(1.8, 1.0), S_c = S_c,
m_t = NULL, m_c = NULL,
kappa0_t = 2.0, nu0_t = 5.0, mu0_t = c(2.0, 1.0), Lambda0_t = L0,
kappa0_c = 2.0, nu0_c = 5.0, mu0_c = c(0.0, 0.0), Lambda0_c = L0,
r = NULL,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 5000L
)
print(round(p_post_niw, 4))
#> R1 R2 R3 R4 R5 R6 R7 R8 R9
#> 0.5116 0.2228 0.0000 0.1320 0.1324 0.0010 0.0002 0.0000 0.0000Posterior predictive probability (future Phase III: ):
set.seed(42)
p_pred <- pbayespostpred2cont(
prob = 'predictive', design = 'controlled', prior = 'vague',
theta_TV1 = NULL, theta_MAV1 = NULL,
theta_TV2 = NULL, theta_MAV2 = NULL,
theta_NULL1 = 0.5, theta_NULL2 = 0.3,
n_t = 20L, n_c = 20L,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = c(1.8, 1.0), S_c = S_c,
m_t = 60L, m_c = 60L,
kappa0_t = NULL, nu0_t = NULL, mu0_t = NULL, Lambda0_t = NULL,
kappa0_c = NULL, nu0_c = NULL, mu0_c = NULL, Lambda0_c = NULL,
r = NULL,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 5000L
)
print(round(p_pred, 4))
#> R1 R2 R3 R4
#> 1 0 0 0
cat(sprintf(
"\nGo region (R1): P = %.4f >= gamma1 (0.80)? %s\n",
p_pred["R1"], ifelse(p_pred["R1"] >= 0.80, "YES", "NO")
))
#>
#> Go region (R1): P = 1.0000 >= gamma1 (0.80)? YES
cat(sprintf(
"NoGo region (R4): P = %.4f >= gamma2 (0.20)? %s\n",
p_pred["R4"], ifelse(p_pred["R4"] >= 0.20, "YES", "NO")
))
#> NoGo region (R4): P = 0.0000 >= gamma2 (0.20)? NO
cat(sprintf("Decision: %s\n",
ifelse(p_pred["R1"] >= 0.80 & p_pred["R4"] < 0.20, "Go",
ifelse(p_pred["R1"] < 0.80 & p_pred["R4"] >= 0.20, "NoGo",
ifelse(p_pred["R1"] >= 0.80 & p_pred["R4"] >= 0.20, "Miss",
"Gray")))
))
#> Decision: Go4.2 Uncontrolled Design
No concurrent control group is enrolled. Under the NIW prior, the hypothetical control distribution is
where
(mu0_c) is the assumed control mean,
(r) scales the covariance relative to the treatment group,
and
is the posterior scale matrix of the treatment group.
set.seed(1)
p_unctrl <- pbayespostpred2cont(
prob = 'posterior', design = 'uncontrolled', prior = 'N-Inv-Wishart',
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = NULL,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = NULL, S_c = NULL,
m_t = NULL, m_c = NULL,
kappa0_t = 2.0, nu0_t = 5.0, mu0_t = c(2.0, 1.0), Lambda0_t = L0,
kappa0_c = NULL, nu0_c = NULL, mu0_c = c(0.0, 0.0), Lambda0_c = NULL,
r = 1.0,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 5000L
)
print(round(p_unctrl, 4))
#> R1 R2 R3 R4 R5 R6 R7 R8 R9
#> 1 0 0 0 0 0 0 0 04.3 External Design (Power Prior)
External data for group
(sample size
,
sample mean
,
sum-of-squares matrix
)
are incorporated via a power prior with weight
.
Both prior = 'vague' and
prior = 'N-Inv-Wishart' are supported.
Vague prior (prior = 'vague')
The posterior parameters after combining the vague power prior with the PoC data are given by Corollary 2 of Huang et al. (2025):
The marginal posterior of is then
S_t <- matrix(c(18.0, 3.6, 3.6, 9.0), 2, 2)
S_c <- matrix(c(16.0, 2.8, 2.8, 8.5), 2, 2)
Se2_ext <- matrix(c(15.0, 2.5, 2.5, 7.5), 2, 2)
set.seed(2)
p_ext_vague <- pbayespostpred2cont(
prob = 'posterior', design = 'external', prior = 'vague',
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = 20L,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = c(1.8, 1.0), S_c = S_c,
m_t = NULL, m_c = NULL,
kappa0_t = NULL, nu0_t = NULL, mu0_t = NULL, Lambda0_t = NULL,
kappa0_c = NULL, nu0_c = NULL, mu0_c = NULL, Lambda0_c = NULL,
r = NULL,
ne_t = NULL, ne_c = 10L, alpha0e_t = NULL, alpha0e_c = 0.5,
bar_ye_t = NULL, bar_ye_c = c(1.5, 0.8), se_t = NULL, se_c = Se2_ext,
nMC = 5000L
)
print(round(p_ext_vague, 4))
#> R1 R2 R3 R4 R5 R6 R7 R8 R9
#> 0.6164 0.1900 0.0002 0.1184 0.0750 0.0000 0.0000 0.0000 0.0000N-Inv-Wishart prior (prior = 'N-Inv-Wishart')
The power prior first combines the initial NIW prior with the discounted external data (Theorem 5 of Huang et al., 2025):
This intermediate NIW result is then updated with the current PoC data by standard conjugate updating (Theorem 6 of Huang et al., 2025):
The marginal posterior of is then
S_t <- matrix(c(18.0, 3.6, 3.6, 9.0), 2, 2)
S_c <- matrix(c(16.0, 2.8, 2.8, 8.5), 2, 2)
L0 <- matrix(c(8.0, 0.0, 0.0, 2.0), 2, 2)
Se2_ext <- matrix(c(15.0, 2.5, 2.5, 7.5), 2, 2)
set.seed(3)
p_ext <- pbayespostpred2cont(
prob = 'posterior', design = 'external', prior = 'N-Inv-Wishart',
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = 20L,
ybar_t = c(3.5, 2.1), S_t = S_t,
ybar_c = c(1.8, 1.0), S_c = S_c,
m_t = NULL, m_c = NULL,
kappa0_t = 2.0, nu0_t = 5.0, mu0_t = c(2.0, 1.0), Lambda0_t = L0,
kappa0_c = 2.0, nu0_c = 5.0, mu0_c = c(0.0, 0.0), Lambda0_c = L0,
r = NULL,
ne_t = NULL, ne_c = 10L, alpha0e_t = NULL, alpha0e_c = 0.5,
bar_ye_t = NULL, bar_ye_c = c(1.5, 0.8), se_t = NULL, se_c = Se2_ext,
nMC = 5000L
)
print(round(p_ext, 4))
#> R1 R2 R3 R4 R5 R6 R7 R8 R9
#> 0.5674 0.2008 0.0000 0.1212 0.1100 0.0002 0.0000 0.0004 0.00005. Operating Characteristics
5.1 Definition
For given true parameters
(mu_t, Sigma_t, mu_c,
Sigma_c), the operating characteristics are estimated by
Monte Carlo over
replicates (nsim). Let
and
for the
-th
simulated dataset. Then:
Here
and
correspond to arguments gamma_go and
gamma_nogo, respectively. A Miss
(
and
simultaneously) indicates an inconsistent threshold configuration and
triggers an error by default (error_if_Miss = TRUE).
5.2 Example: Controlled Design, Posterior Probability
Treatment effect scenarios are defined over a two-dimensional grid of
while fixing
.
The full factorial combination of unique
and
values triggers the tile plot in plot(), with colour
intensity encoding
:
Sigma <- matrix(c(4.0, 0.8, 0.8, 1.0), 2, 2)
mu_t1_seq <- seq(0.0, 3.5, by = 0.5)
mu_t2_seq <- seq(0.0, 2.1, by = 0.3)
n_scen <- length(mu_t1_seq) * length(mu_t2_seq)
oc_ctrl <- pbayesdecisionprob2cont(
nsim = 100L, prob = 'posterior', design = 'controlled',
prior = 'vague',
GoRegions = 1L, NoGoRegions = 9L,
gamma_go = 0.80, gamma_nogo = 0.20,
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
n_t = 20L, n_c = 20L, m_t = NULL, m_c = NULL,
mu_t = cbind(rep(mu_t1_seq, times = length(mu_t2_seq)),
rep(mu_t2_seq, each = length(mu_t1_seq))),
Sigma_t = Sigma,
mu_c = matrix(0, nrow = n_scen, ncol = 2),
Sigma_c = Sigma,
kappa0_t = NULL, nu0_t = NULL, mu0_t = NULL, Lambda0_t = NULL,
kappa0_c = NULL, nu0_c = NULL, mu0_c = NULL, Lambda0_c = NULL,
r = NULL,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 500L, CalcMethod = 'MC',
error_if_Miss = TRUE, Gray_inc_Miss = FALSE, seed = 42L
)
print(oc_ctrl)
#> Go/NoGo/Gray Decision Probabilities (Two Continuous Endpoints)
#> ----------------------------------------------------------------
#> Probability type : posterior
#> Design : controlled
#> Prior : vague
#> Method : NULL
#> Simulations : nsim = 100
#> MC draws : nMC = 500
#> Seed : 42
#> Threshold(s) : TV1 = 1.5, MAV1 = 0.5
#> TV2 = 1, MAV2 = 0.3
#> Go threshold : gamma_go = 0.8
#> NoGo threshold : gamma_nogo = 0.2
#> Go regions : {1}
#> NoGo regions : {9}
#> Sample size : n_t = 20, n_c = 20
#> True cov (treat.): Sigma_t = [4, 0.8; 0.8, 1]
#> True cov (cont.) : Sigma_c = [4, 0.8; 0.8, 1]
#> Miss handling : error_if_Miss = TRUE, Gray_inc_Miss = FALSE
#> ----------------------------------------------------------------
#> mu_t1 mu_t2 mu_c1 mu_c2 Go Gray NoGo
#> 0.0 0.0 0 0 0.0000 0.0800 0.9200
#> 0.5 0.0 0 0 0.0000 0.2100 0.7900
#> 1.0 0.0 0 0 0.0000 0.5900 0.4100
#> 1.5 0.0 0 0 0.0000 0.8600 0.1400
#> 2.0 0.0 0 0 0.0000 0.9400 0.0600
#> 2.5 0.0 0 0 0.0000 0.9900 0.0100
#> 3.0 0.0 0 0 0.0000 1.0000 0.0000
#> 3.5 0.0 0 0 0.0000 1.0000 0.0000
#> 0.0 0.3 0 0 0.0000 0.2600 0.7400
#> 0.5 0.3 0 0 0.0000 0.4000 0.6000
#> 1.0 0.3 0 0 0.0000 0.6800 0.3200
#> 1.5 0.3 0 0 0.0000 0.8700 0.1300
#> 2.0 0.3 0 0 0.0000 0.9400 0.0600
#> 2.5 0.3 0 0 0.0000 0.9900 0.0100
#> 3.0 0.3 0 0 0.0000 1.0000 0.0000
#> 3.5 0.3 0 0 0.0000 1.0000 0.0000
#> 0.0 0.6 0 0 0.0000 0.6200 0.3800
#> 0.5 0.6 0 0 0.0000 0.6600 0.3400
#> 1.0 0.6 0 0 0.0000 0.7900 0.2100
#> 1.5 0.6 0 0 0.0000 0.9300 0.0700
#> 2.0 0.6 0 0 0.0200 0.9400 0.0400
#> 2.5 0.6 0 0 0.0300 0.9600 0.0100
#> 3.0 0.6 0 0 0.0300 0.9700 0.0000
#> 3.5 0.6 0 0 0.0300 0.9700 0.0000
#> 0.0 0.9 0 0 0.0000 0.8700 0.1300
#> 0.5 0.9 0 0 0.0000 0.9000 0.1000
#> 1.0 0.9 0 0 0.0100 0.9200 0.0700
#> 1.5 0.9 0 0 0.0400 0.9000 0.0600
#> 2.0 0.9 0 0 0.0400 0.9300 0.0300
#> 2.5 0.9 0 0 0.0600 0.9400 0.0000
#> 3.0 0.9 0 0 0.1100 0.8900 0.0000
#> 3.5 0.9 0 0 0.1400 0.8600 0.0000
#> 0.0 1.2 0 0 0.0000 0.9500 0.0500
#> 0.5 1.2 0 0 0.0000 0.9500 0.0500
#> 1.0 1.2 0 0 0.0100 0.9700 0.0200
#> 1.5 1.2 0 0 0.0500 0.9300 0.0200
#> 2.0 1.2 0 0 0.1200 0.8800 0.0000
#> 2.5 1.2 0 0 0.3000 0.7000 0.0000
#> 3.0 1.2 0 0 0.4000 0.6000 0.0000
#> 3.5 1.2 0 0 0.3900 0.6100 0.0000
#> 0.0 1.5 0 0 0.0000 1.0000 0.0000
#> 0.5 1.5 0 0 0.0000 1.0000 0.0000
#> 1.0 1.5 0 0 0.0300 0.9700 0.0000
#> 1.5 1.5 0 0 0.0800 0.9200 0.0000
#> 2.0 1.5 0 0 0.2700 0.7300 0.0000
#> 2.5 1.5 0 0 0.5900 0.4100 0.0000
#> 3.0 1.5 0 0 0.7400 0.2600 0.0000
#> 3.5 1.5 0 0 0.7700 0.2300 0.0000
#> 0.0 1.8 0 0 0.0000 1.0000 0.0000
#> 0.5 1.8 0 0 0.0000 1.0000 0.0000
#> 1.0 1.8 0 0 0.0300 0.9700 0.0000
#> 1.5 1.8 0 0 0.1000 0.9000 0.0000
#> 2.0 1.8 0 0 0.4100 0.5900 0.0000
#> 2.5 1.8 0 0 0.7600 0.2400 0.0000
#> 3.0 1.8 0 0 0.8800 0.1200 0.0000
#> 3.5 1.8 0 0 0.9300 0.0700 0.0000
#> 0.0 2.1 0 0 0.0000 1.0000 0.0000
#> 0.5 2.1 0 0 0.0000 1.0000 0.0000
#> 1.0 2.1 0 0 0.0300 0.9700 0.0000
#> 1.5 2.1 0 0 0.1000 0.9000 0.0000
#> 2.0 2.1 0 0 0.3600 0.6400 0.0000
#> 2.5 2.1 0 0 0.8000 0.2000 0.0000
#> 3.0 2.1 0 0 0.9200 0.0800 0.0000
#> 3.5 2.1 0 0 0.9700 0.0300 0.0000
#> ----------------------------------------------------------------
plot(oc_ctrl, base_size = 20)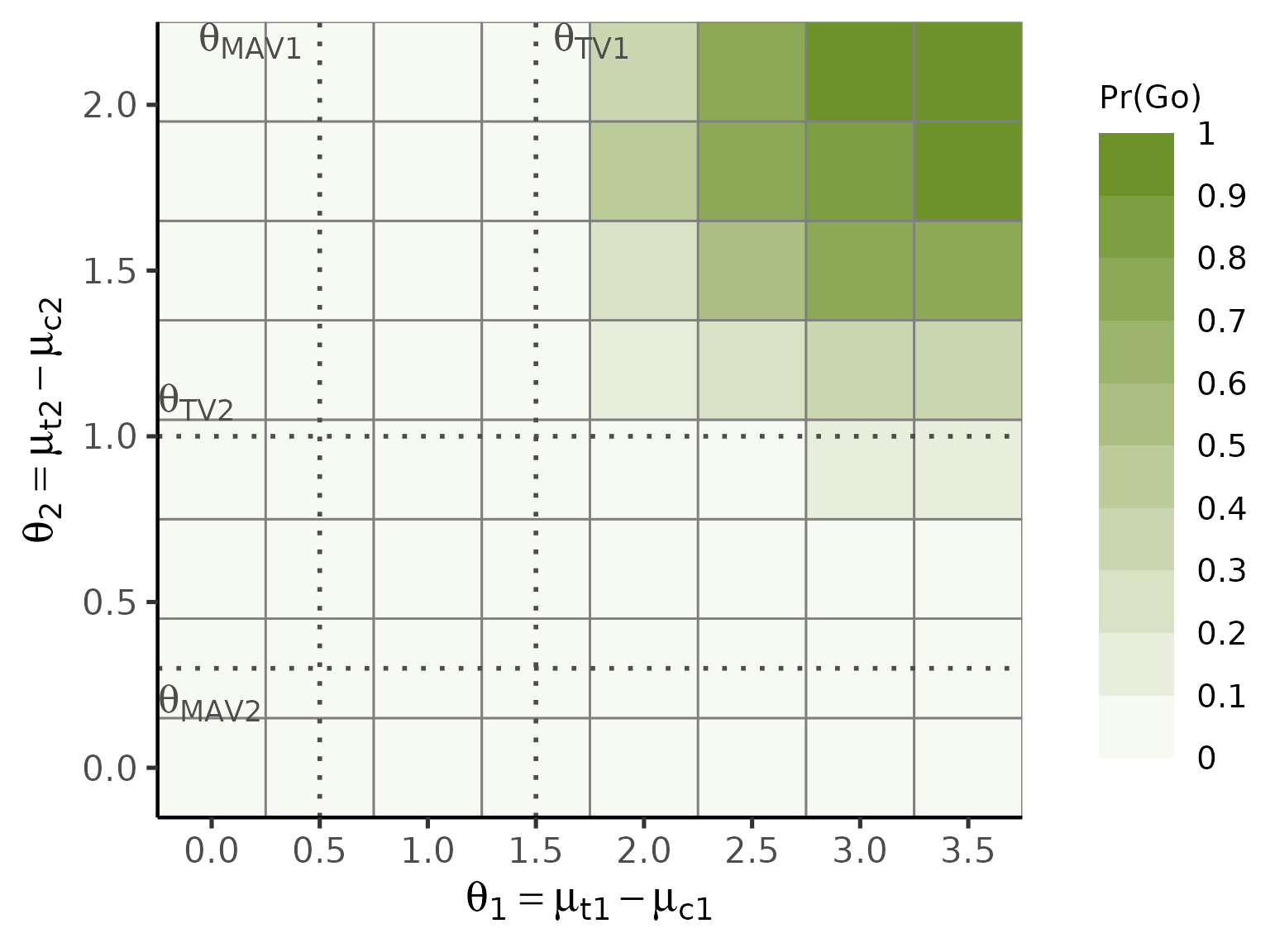
The same function supports design = 'uncontrolled' (with
hypothetical control mean mu0_c and scale factor
r) and design = 'external' (with power prior
arguments ne_c, bar_ye_c, se_c,
alpha0e_c). The argument prob = 'predictive'
switches to predictive probability (with future sample sizes
m_t, m_c and null thresholds
theta_NULL1, theta_NULL2). The function
signature and output format are identical across all combinations.
6. Optimal Threshold Search
6.1 Objective and Algorithm
Because there are two decision thresholds
,
the optimal search is performed over a two-dimensional grid
(arguments gamma_go_grid and gamma_nogo_grid).
The two thresholds are calibrated independently under separate
scenarios:
-
:
calibrated under the Go-calibration scenario (
mu_t_go,Sigma_t_go,mu_c_go,Sigma_c_go; typically the null scenario ), so that . -
:
calibrated under the NoGo-calibration scenario (
mu_t_nogo,Sigma_t_nogo,mu_c_nogo,Sigma_c_nogo; typically the alternative scenario), so that .
getgamma2cont() uses a three-stage strategy:
Stage 1 (Simulate and precompute):
bivariate datasets are generated separately from each calibration
scenario. For each replicate
,
pbayespostpred2cont() is called once in vectorised mode to
return the full region probability vector. The marginal Go and NoGo
probabilities per replicate are:
Stage 2 (Gamma sweep): For each candidate (under the Go-calibration scenario) and each (under the NoGo-calibration scenario):
The results are stored as vectors PrGo_grid and
PrNoGo_grid indexed by gamma_go_grid and
gamma_nogo_grid, respectively.
Stage 3 (Optimal selection): is the smallest grid value satisfying ; is the smallest grid value satisfying .
6.2 Example: Controlled Design, Posterior Probability
Go-calibration scenario: , (effect at MAV boundary – above which Go is not yet warranted, so is required). NoGo-calibration scenario: , (effect between MAV and TV – above which NoGo should be unlikely).
res_gamma <- getgamma2cont(
nsim = 500L, prob = 'posterior', design = 'controlled',
prior = 'vague',
GoRegions = 1L, NoGoRegions = 9L,
mu_t_go = c(0.5, 0.3), Sigma_t_go = Sigma,
mu_c_go = c(0.0, 0.0), Sigma_c_go = Sigma,
mu_t_nogo = c(1.0, 0.6), Sigma_t_nogo = Sigma,
mu_c_nogo = c(0.0, 0.0), Sigma_c_nogo = Sigma,
target_go = 0.05, target_nogo = 0.20,
n_t = 20L, n_c = 20L,
theta_TV1 = 1.5, theta_MAV1 = 0.5,
theta_TV2 = 1.0, theta_MAV2 = 0.3,
theta_NULL1 = NULL, theta_NULL2 = NULL,
m_t = NULL, m_c = NULL,
kappa0_t = NULL, nu0_t = NULL, mu0_t = NULL, Lambda0_t = NULL,
kappa0_c = NULL, nu0_c = NULL, mu0_c = NULL, Lambda0_c = NULL,
r = NULL,
ne_t = NULL, ne_c = NULL, alpha0e_t = NULL, alpha0e_c = NULL,
bar_ye_t = NULL, bar_ye_c = NULL, se_t = NULL, se_c = NULL,
nMC = 500L, CalcMethod = 'MC',
gamma_go_grid = seq(0.05, 0.95, by = 0.05),
gamma_nogo_grid = seq(0.05, 0.95, by = 0.05),
seed = 42L
)
plot(res_gamma, base_size = 20)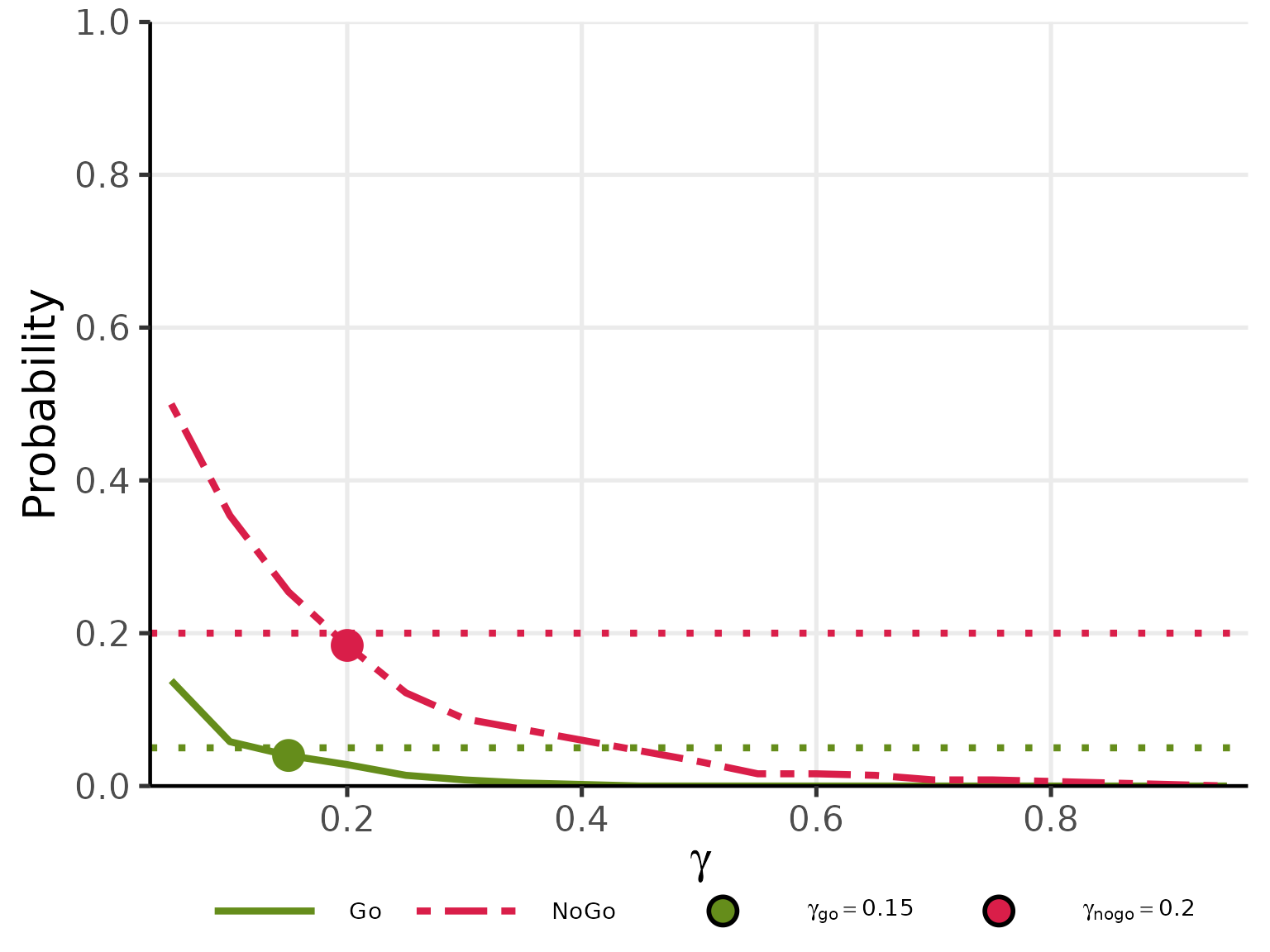
The same function supports design = 'uncontrolled' (with
mu0_c and r) and
design = 'external' (with power prior arguments
ne_c, bar_ye_c, se_c,
alpha0e_c). Setting prob = 'predictive'
switches to predictive probability calibration (with
theta_NULL1, theta_NULL2, m_t,
m_c). The function signature is identical across all
combinations.
7. Summary
This vignette covered two continuous endpoint analysis in BayesianQDM:
- Bayesian model: NIW conjugate posterior with explicit update formulae for , , , and ; bivariate non-standardised marginal posterior.
- Nine-region grid for posterior probability and four-region grid for predictive probability; typical choice Go = R1, NoGo = R9 (posterior) or R4 (predictive).
- Two computation methods: MC (vectorised pre-generated draws with
Cholesky reuse) and MM (Mahalanobis-distance fourth-moment matching via
Yamaguchi et al. (2025), Theorem 3, +
mvtnorm::pmvtfor joint probability; requires ). - Three study designs: controlled, uncontrolled ( and fixed), external design (vague power prior follows Corollary 2 of Huang et al. (2025); NIW power prior follows Theorems 5–6).
- Operating characteristics: Monte Carlo estimation with vectorised region probability computation.
- Optimal threshold search: three-stage strategy in
getgamma2cont()calibrating and independently under separate Go- and NoGo-calibration scenarios, with calibration curves visualised viaplot()and optimal thresholds marked at the target probability levels ( under null, under alternative).
See vignette("two-binary") for the analogous two binary
endpoint analysis.
